-Search query
-Search result
Showing 1 - 50 of 291 items for (author: choi & e)
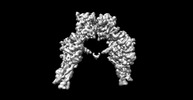
EMDB-41877: 
Cryo-EM structure of long form insulin receptor (IR-B) in the apo state
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E
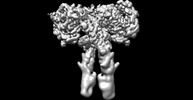
EMDB-41878: 
Cryo-EM structure of long form insulin receptor (IR-B) with four IGF2 bound, symmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

EMDB-41880: 
Cryo-EM structure of long form insulin receptor (IR-B) with three IGF2 bound, asymmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E
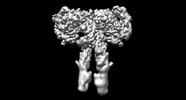
EMDB-43279: 
Cryo-EM structure of short form insulin receptor (IR-A) with four IGF2 bound, symmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

EMDB-43280: 
Cryo-EM structure of short form insulin receptor (IR-A) with three IGF2 bound, asymmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8u4b: 
Cryo-EM structure of long form insulin receptor (IR-B) in the apo state
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8u4c: 
Cryo-EM structure of long form insulin receptor (IR-B) with four IGF2 bound, symmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8u4e: 
Cryo-EM structure of long form insulin receptor (IR-B) with three IGF2 bound, asymmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8vjb: 
Cryo-EM structure of short form insulin receptor (IR-A) with four IGF2 bound, symmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

PDB-8vjc: 
Cryo-EM structure of short form insulin receptor (IR-A) with three IGF2 bound, asymmetric conformation.
Method: single particle / : An W, Hall C, Li J, Huang A, Wu J, Park J, Bai XC, Choi E

EMDB-29659: 
CryoEM structure of nuclear GAPDH under 8h Oxidative Stress
Method: single particle / : Choi WY, Wu H, Cheng YF, Manglik A

EMDB-29660: 
CryoEM structure of cytosolic GAPDH under 8h Oxidative Stress
Method: single particle / : Choi WY, Wu H, Cheng YF, Manglik A

EMDB-29661: 
CryoEM structure of cytosolic GAPDH under 8h Oxidative Stress, class2
Method: single particle / : Choi WY, Wu H, Cheng YF, Manglik A

EMDB-29662: 
CryoEM structure of nuclear GAPDH under 24h Oxidative Stress
Method: single particle / : Choi WY, Wu H, Cheng YF, Manglik A

EMDB-29663: 
CryoEM structure of cytoplasmic GAPDH under 24h Oxidative Stress
Method: single particle / : Choi WY, Wu H, Cheng YF, Manglik A

EMDB-29664: 
CryoEM structure of wild-type GAPDH
Method: single particle / : Choi WY, Wu H, Cheng YF, Manglik A

PDB-8g12: 
CryoEM structure of nuclear GAPDH under 8h Oxidative Stress
Method: single particle / : Choi WY, Wu H, Cheng YF

PDB-8g13: 
CryoEM structure of cytosolic GAPDH under 8h Oxidative Stress
Method: single particle / : Choi WY, Wu H, Cheng YF

PDB-8g14: 
CryoEM structure of cytosolic GAPDH under 8h Oxidative Stress, class2
Method: single particle / : Choi WY, Wu H, Cheng YF

PDB-8g15: 
CryoEM structure of nuclear GAPDH under 24h Oxidative Stress
Method: single particle / : Choi WY, Wu H, Cheng YF

PDB-8g16: 
CryoEM structure of cytoplasmic GAPDH under 24h Oxidative Stress
Method: single particle / : Choi WY, Wu H, Cheng YF

EMDB-35010: 
Human Consensus Olfactory Receptor OR52c in Complex with Octanoic Acid (OCA) and G Protein
Method: single particle / : Choi CW, Bae J, Choi HJ, Kim J

EMDB-35770: 
Human Consensus Olfactory Receptor OR52c in Complex with Octanoic Acid (OCA) and G Protein (Consensus map)
Method: single particle / : Choi CW, Bae J, Choi HJ

EMDB-35772: 
Human Consensus Olfactory Receptor OR52c in Complex with Octanoic Acid (OCA) and G Protein (Receptor-focused map)
Method: single particle / : Choi CW, Bae J, Choi HJ

EMDB-35773: 
Human Consensus Olfactory Receptor OR52c in Complex with Octanoic Acid (OCA) and G Protein (G protein-focused map)
Method: single particle / : Choi CW, Bae J, Choi HJ

EMDB-35971: 
Human Consensus Olfactory Receptor OR52c in apo state, OR52c-bRIL
Method: single particle / : Choi CW, Bae J, Choi HJ, Kim J
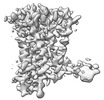
EMDB-37336: 
Human Consensus Olfactory Receptor OR52c in apo state, OR52c only
Method: single particle / : Choi CW, Bae J, Choi HJ, Kim J

PDB-8hti: 
Human Consensus Olfactory Receptor OR52c in Complex with Octanoic Acid (OCA) and G Protein
Method: single particle / : Choi CW, Bae J, Choi HJ, Kim J

PDB-8j46: 
Human Consensus Olfactory Receptor OR52c in apo state, OR52c-bRIL
Method: single particle / : Choi CW, Bae J, Choi HJ, Kim J

PDB-8w77: 
Human Consensus Olfactory Receptor OR52c in apo state, OR52c only
Method: single particle / : Choi CW, Bae J, Choi HJ, Kim J

EMDB-40699: 
SARS-CoV-2 replication-transcription complex bound to nsp9 and UMPCPP, as a pre-catalytic NMPylation intermediate
Method: single particle / : Small GI, Darst SA, Campbell EA

EMDB-40707: 
SARS-CoV-2 replication-transcription complex bound to RNA-nsp9, as a noncatalytic RNA-nsp9 binding mode
Method: single particle / : Small GI, Darst SA, Campbell EA

EMDB-40708: 
SARS-CoV-2 replication-transcription complex bound to RNA-nsp9 and GDP-betaS, as a pre-catalytic deRNAylation/mRNA capping intermediate
Method: single particle / : Small GI, Darst SA, Campbell EA

PDB-8sq9: 
SARS-CoV-2 replication-transcription complex bound to nsp9 and UMPCPP, as a pre-catalytic NMPylation intermediate
Method: single particle / : Small GI, Darst SA, Campbell EA

PDB-8sqj: 
SARS-CoV-2 replication-transcription complex bound to RNA-nsp9, as a noncatalytic RNA-nsp9 binding mode
Method: single particle / : Small GI, Darst SA, Campbell EA

PDB-8sqk: 
SARS-CoV-2 replication-transcription complex bound to RNA-nsp9 and GDP-betaS, as a pre-catalytic deRNAylation/mRNA capping intermediate
Method: single particle / : Small GI, Darst SA, Campbell EA

EMDB-28823: 
Phi-29 partially-expanded fiberless prohead
Method: single particle / : Woodson ME, Morais MC, Scott SD, Choi KH, Jardine PJ, Zhang W

PDB-8f2n: 
Phi-29 partially-expanded fiberless prohead
Method: single particle / : Woodson ME, Morais MC, Scott SD, Choi KH, Jardine PJ, Zhang W

EMDB-28820: 
phi-29 prohead MCP gp8 penton maturation intermediate, with associated scaffold gp7 tetramer
Method: single particle / : Woodson ME, Morais MC, Jardine PJ, Zhang W, Prokhorov NS

EMDB-28821: 
bacteriophage phi-29 MCP gp-8 penton in intermediate maturation state, with associated scaffold gp7 dimer
Method: single particle / : Woodson ME, Morais MC, Prokhorov NS, Zhang W, Jardine PJ

EMDB-28822: 
Phi-29 scaffolding protein bound to intermediate-state MCP
Method: single particle / : Woodson ME, Morais MC, Jardine PJ, Scott SD

EMDB-28824: 
Phi-29 expanded, DNA-packaged fiberless prohead
Method: single particle / : Woodson ME, Morais MC, Jardine PJ, Zhang W

EMDB-29013: 
96nm doublet microtubule repeat from wild type mouse sperm
Method: subtomogram averaging / : Hwang JY, Chai P, Nawaz S

EMDB-28606: 
96nm doublet microtubule repeat from LRRC23-KO mouse sperm
Method: subtomogram averaging / : Hwang JY, Chai P, Nawaz S

EMDB-36937: 
post-occluded structure of human ABCB6 W546A mutant (ADP/VO4-bound)
Method: single particle / : Jin MS, Lee SS, Park JG, Jang E, Choi SH, Kim S, Kim JW

EMDB-36938: 
Outward-facing structure of human ABCB6 W546A mutant (ADP/VO4-bound)
Method: single particle / : Jin MS, Lee SS, Park JG, Jang E, Choi SH, Kim S, Kim JW

PDB-8k7b: 
post-occluded structure of human ABCB6 W546A mutant (ADP/VO4-bound)
Method: single particle / : Jin MS, Lee SS, Park JG, Jang E, Choi SH, Kim S, Kim JW

PDB-8k7c: 
Outward-facing structure of human ABCB6 W546A mutant (ADP/VO4-bound)
Method: single particle / : Jin MS, Lee SS, Park JG, Jang E, Choi SH, Kim S, Kim JW

EMDB-40305: 
Cryo-EM structure of insulin amyloid-like fibril that is composed of two antiparallel protofilaments
Method: single particle / : Wang LW, Hall C, Uchikawa E, Chen DL, Choi E, Zhang XW, Bai XC
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
